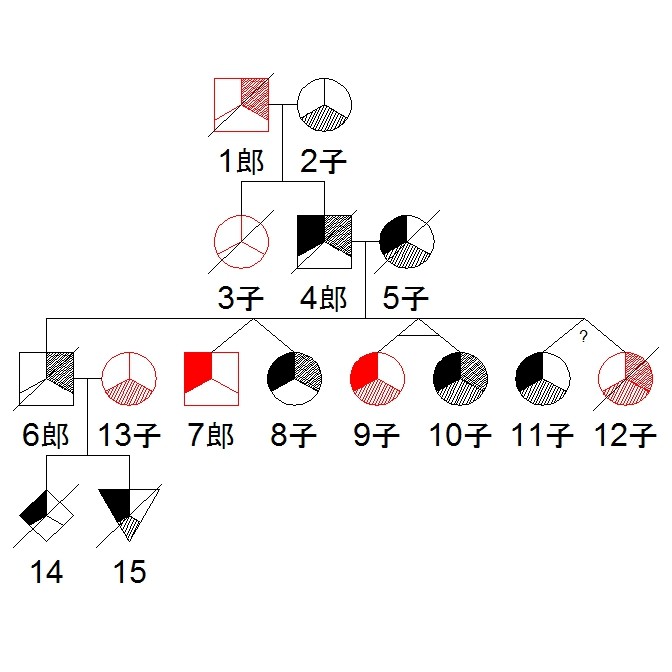
library(kinship2)
id <- 1:15
sex <- c(1,2,2,1,2,1,1,2,2,2,2,2,2,3,4)
lab <- id
lab[which(sex==1)] <- paste(lab[which(sex==1)],"郎",sep="")
lab[which(sex==2)] <- paste(lab[which(sex==2)],"子",sep="")
num.phenotype <- 3
affected <- matrix(sample(0:1,length(id)*num.phenotype,replace=TRUE),ncol=num.phenotype)
status <- sample(0:1,length(id),replace=TRUE)
relation <- matrix(c(6,7,2,6,8,2,7,8,2,9,10,1,11,12,3,6,13,4),ncol=3,byrow=TRUE)
dadid <- c(0,0,1,1,0,4,4,4,4,4,4,4,0,6,6)
momid <- c(0,0,2,2,0,5,5,5,5,5,5,5,0,13,13)
my.ped <- pedigree(id,dadid,momid,sex,affected,status,relation)
avail <- sample(1:2,length(id),replace=TRUE)
plot(my.ped,id=lab,col=avail,cex=2)
